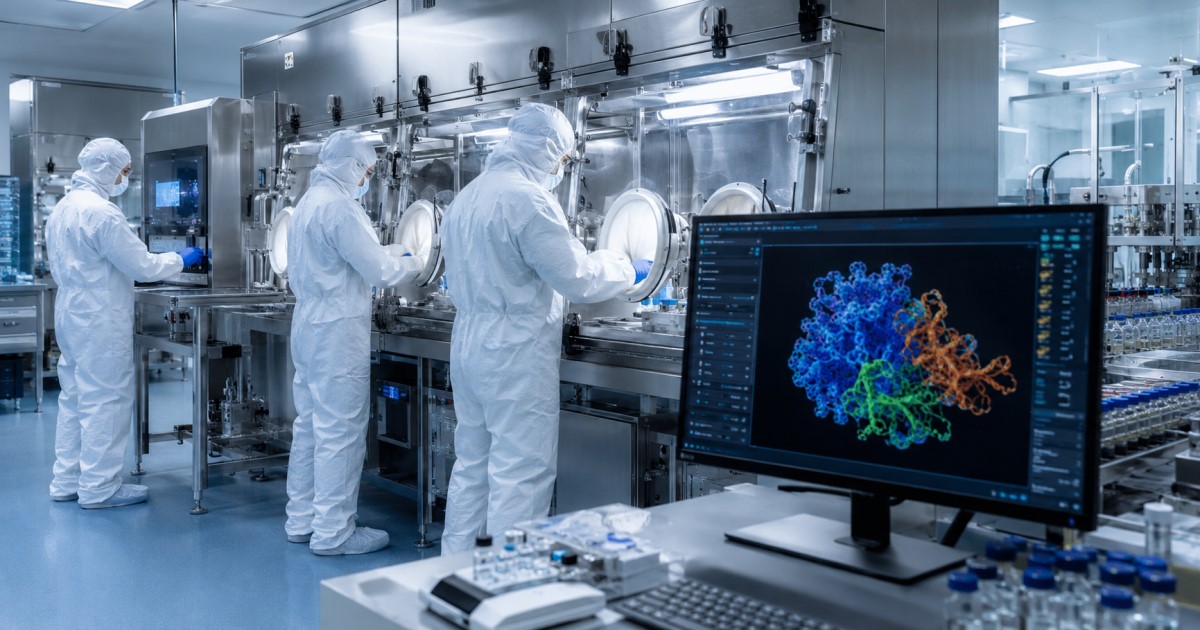

AI-designed drugs are reaching humans
This moves AI drug discovery beyond software prediction into early human translational evidence.
Machine-Readable Science • ISSN 3054-3991

DOI: 10.62487/saimsara62d4eb26

This moves AI drug discovery beyond software prediction into early human translational evidence.
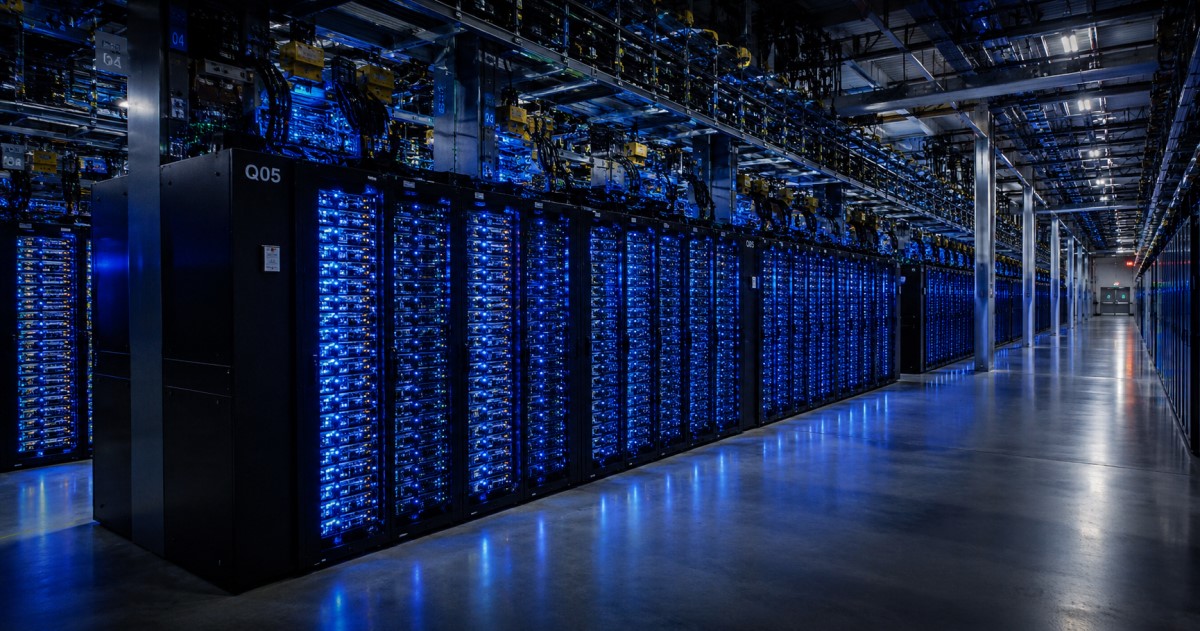

The discovery bottleneck is shifting from “can we search this space?” to “which hits are biologically real?”
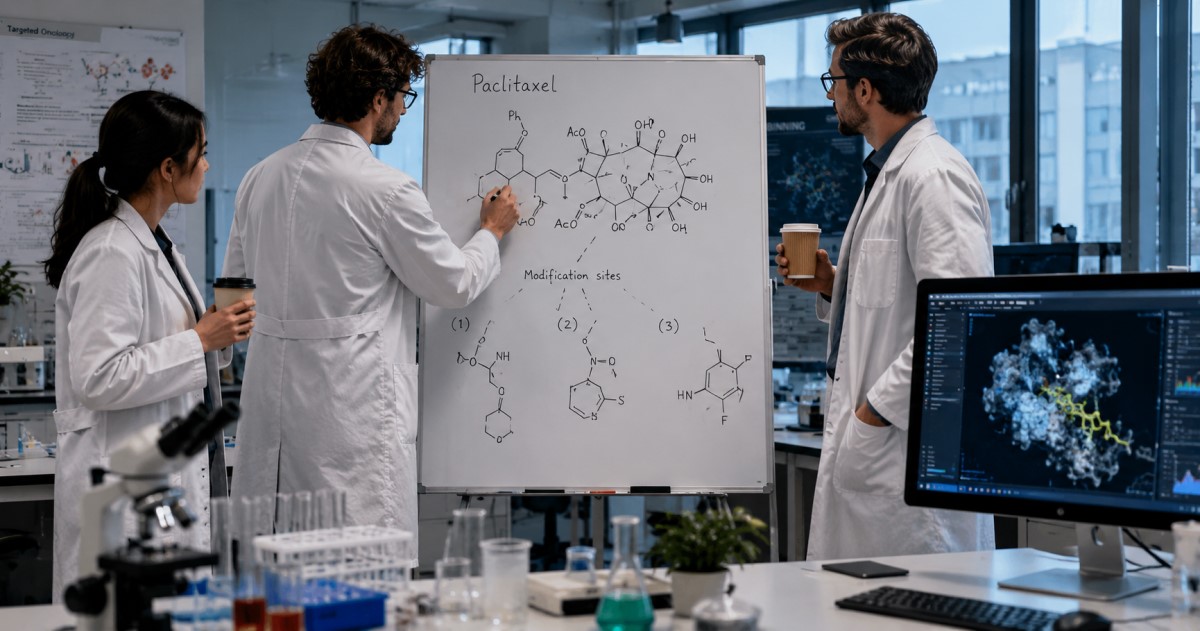
This is the safeguard signal: powerful AI still needs expert review, realistic validation, and human-in-the-loop decisions.
Unlock the full evidence map
Full paper access includes the complete human-readable review, figures, reference index, PDF export, and machine-readable Evidence JSON download.
Evidence JSON can also be purchased separately if you only need the LLM-ready object for agent, AI, or RAG workflows.
Institutional or library access? Sign in with your institution email to open all available SAIMSARA papers under your institution access arrangement.
Need a SAIMSARA review on your own topic? ☸️Request.
Unlock the full paper to view the complete Reference Index.